|
subtilis genome harbors two 6S RNA homologs, termed 6S-1 and 6S-2 ( 11, 12). Synthesis of these short transcripts leads to dissociation of 6S RNA–RNAP complexes. This highly conserved secondary structure is essential for the ability of 6S RNA to form stable complexes with Eσ 70 and to serve as a template for the synthesis of pRNAs ( 9, 10). 6S RNA is primarily helical, with a large single-stranded loop in the center. In Escherichia coli, this RNA acts by complex formation with the σ 70 housekeeping RNA polymerase (RNAP) holoenzyme (Eσ 70) and accumulates during stationary phase. 6S RNA, ∼200 nt in size, has been identified as a growth-phase dependent riboregulator of bacterial transcription ( 8). We were faced with the limitations of standard northern blot protocols when trying to detect small so-called product RNAs (pRNAs, ∼14 nt in length) in total RNA extracts from Bacillus subtilis pRNAs are short transcripts synthesized by bacterial RNA polymerases in an RNA-dependent RNA polymerization reaction using 6S RNA as template. Furthermore, the number and positions of UV-crosslinked bases vary depending on the particular RNA sequence and the applied UV crosslinking conditions (length of irradiation, UV light intensity), making UV crosslinking a process that is hard to control ( 6, 7). The likely reason is that UV crosslinking to the free amines of nylon membranes occurs via the bases (primarily uracil) which, in contrast to immobilization via the phosphate, sterically blocks accessibility of the immobilized nucleic acid for hybridization, particularly in the case of small RNAs owing to their limited interaction surface. Relative to the standard UV crosslinking to nylon membranes, this reportedly 5′-end-selective attachment enhanced the detection of RNAs <40 nt 25- to 50-fold. Recently, Hamilton and co-workers ( 6, 7) reported the use of 1-ethyl-3-(3-dimethylaminopropyl)-carbodiimide (EDC) to covalently couple small RNAs via their 5′-phosphates to amino groups at the surface of nylon membranes. The northern blot has even gained importance owing to the growing number of small RNA candidates identified by high-throughput and bioinformatic methods, which need to be verified experimentally. Despite these advances, the classic northern blot technique has remained irreplaceable, since the approach allows the researcher to visualize and roughly quantify cellular levels of RNAs and their processing intermediates relative to endogenous RNA standards (e.g. This includes the use of LNA probes in hybridization approaches to increase sensitivity ( 1, 2), development of the looped-primer RT–PCR technique to quantify mature miRNAs ( 3), as well as high-throughput methods such as RNomics to generate cellular RNA libraries for deep sequencing ( 4) or miRNA profiling arrays ( 5). This entire northern blotting protocol takes similar to 4 d to complete.The discovery of small regulatory RNAs in all domains of life and their multi-faceted key roles in cell biology has propelled advances in technologies for their analysis. Despite the development of newer methods, such as real-time PCR, nuclease protection assays and microarrays, northern blot analysis is still a standard technique used in the detection and quantification of mRNA levels because it allows a direct comparison of the mRNA abundance between samples on a single membrane. Specific sequences in the RNA are detected on the membrane by molecular hybridization with radiolabeled nucleic acid probes. RNA molecules are separated by gel electrophoresis and are subsequently transferred to a porous membrane by capillary action. 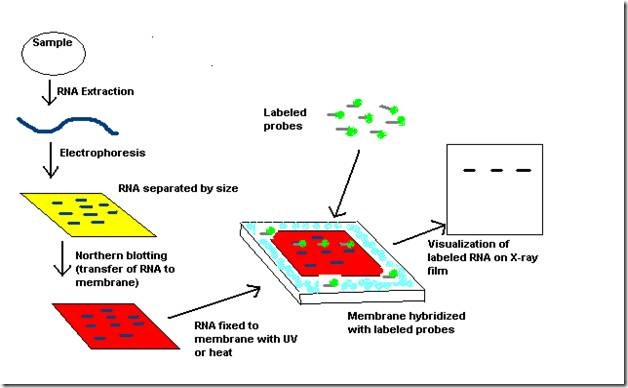
In this protocol, we describe how northern blot analysis is used to identify gene expression patterns at the RNA level in human cancer cells as well as in cancerous and normal tissues. Investigation of gene expression significantly contributes to our knowledge of the regulation and function of genes in many areas of biology. Acibadem Mehmet Ali Aydinlar University Affiliated: No.Journal Indexes: Science Citation Index Expanded (SCI-EXPANDED), Scopus.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
